I attended the inaugural openEHR meetings in China last week - this is how I introduced openEHR…
Bridging the interop chasms
The ultimate PHR?
I've been interested in the notion of a Personal Health Record for a long time. I was involved in the development of HotHealth, which launched at the end of 2000, a not-so-auspicious year, given the dot com crash! By the time HotHealth was completed , all the potential competitors identified in the pre-market environmental scan were defunct. It certainly wasn't easy to get any traction for HotHealth take-up and yet only recently it has been retired. For a couple of years it was successfully used at the Royal Children's Hospital, cut down and re-branded as BetterDiabetes to support teenagers self-manage their diabetes and communicate with their clinicians, but it wasn't sustained.
This is not an uncommon story for PHRs. It is somewhat comforting to see that the course of those such HealthVault and GoogleHealth have also not been smooth and fabulously successful :-)
Why is the PHR so hard?
In recent years I participated in the development of the ISO Technical Report 14292:2012: Personal health records -- Definition, scope and context. In this my major contribution seemed to be introducing the idea of a health information continuum.
However in the past year or so, my notion of an ideal PHR has moved on a little further again. It has arisen on the premise of a health record platform in which standardised health information persists independently of any one software application and can be accessed by any compliant applications, whether consumer- or clinician-focused. And the record of health information can be contributed to by any number of compliant systems - whether a clinical system, a PHR or smartphone app. The focus is on the data, the health record itself; not the applications. You will have seen a number of my previous posts, including here & here!
So, in this kind of new health data utopia, imagine if all my weights were automatically uploaded to my Weight app on my smartphone wirelessly each morning. Over time I could graph this and track my BMI etc. Useful stuff, and this can be done now - but only into dead-end silos of data within a given app.
And what if a new fandangled weight management application came along that I liked better - perhaps it provided more support to help me lose weight. And I want to lose weight. So I add the new app to my smartphone and, hey presto, it can immediately access all my previous weights - all because the data structure in both apps is identical. Thus the data can be unambiguously understood and computed upon within the second app without any data manipulation. Pretty cool. No more data silos; no more data loss. Simply delete the first app from the system, and elect to keep the data within my smartphone health record.
And as I add apps that suit my lifestyle, health needs, and fitness goals etc, I'm gradually accumulating important health information that is probably not available anywhere else. And consider that only I actually know what medicines I'm taking, including over the counter and herbals. The notion of a current medication list is really not in the remit of any clinician, but the motivated consumer! And so if I add an app to start to manage my medications or immunisations this data could be also used across in yet another compliant chronic disease support app for my diabetes or asthma or...
I can gradually build up a record of health information that is useful to me to manage my health, and that is also potentially useful to share with my healthcare providers.
Do you see the difference to current PHR systems?
I can choose apps that are 'best of breed' and applicable to my need or interest.
I'm not locked in to any one app, a mega app that contains stuff I don't want and will never use, with all the overheads and lack of flexibility.
I can 'plug & play' apps into my health record, able to change my mind if I find features, a user interface or workflow that I like better.
And yet the data remains ready for future use and potentially for sharing with my healthcare providers, if and when I choose. How cool is that?
Keep in mind that if those data structures were the same as being used by my clinician systems, then there is also potential for me to receive data from my clinicians and incorporate it into my PHR; similarly there is also potential for me to send data to my clinician and give them the choice of incorporating this into their systems - maybe my blood glucose records directly obtained from my glucometer, my weight measurements, etc. Maybe, one day, even MY current medicine list!
In this proposed flexible data environment we are avoiding the 'one size fits all', behemoth approach, which doesn't seem to have worked well in many situations, both clinical systems or personal health records. Best of all the data is preserved in the non-proprietary, shared format - the beginnings of a universal health record or, at least, a health record platform fully supporting data exchange.
What do you think?
"We have the capability!"
We have had the technology for a purpose-built openEHR-compliant 'plug & play' platform for some time; standalone applications have been built, but just recently it appears that the practical reality of an multi-application platform is also about to happen. "We have the technology. We have the capability..." Stay tuned.
...Reminds me of my 1970's hero, Steve Austin, the Six Million Dollar Man. With apologies to Steve:
"We have the technology. We have the capability to make the world's first universal health platform. openEHR will be that platform. Better than ever before. Robust health data...application independent...semantically interoperable!"
[youtube http://www.youtube.com/watch?v=K7zNY0I5JNI&w=480&h=360]
Preserving health data integrity
How valuable do we really think health data is? How seriously do we take our responsibility to preserve the integrity of our health data? Probably not nearly as much as we should.
Consider the current situation of most clinicians or organisations when purchasing a clinical EHR system. What do they look for? Many possible answers are obvious, but there is one question that I suspect very few are asking. How many consider what data they will be able to export and convert to another format, preserving the current data integrity, at the end of the typical 5-10 year life span of the application? Am I wrong if I suggest it is not many at all?
Despite all the effort that we clinicians put into entering detailed data to create a quality health record we don't seem to often consider the " What next?" scenario. How much and precisely what data will we be able to safely extract, export, transfer or convert into the next, inevitable, clinical system? Ironically, we are simultaneously well aware that clinical systems have a limited technical life span.
Any and all of the health data in a health record is an incredibly valuable asset to the holder, to the patient (if these are not the same entities) and to those downstream with whom we may share it in the future - in terms of $$ invested; manpower resources used to capture, store, classify, update and maintain it; and not least the future value that comes from appropriate and safe clinical decisions being made upon the integrity of existing EHR data.
Yet we don't seem to consider it much... yet. However, as more clinicians are creating increasing amounts of isolated pockets of health data, we should be thinking about it very hard.
Every time we change systems we put our health data at risk - risk of absolute data loss and risk of possible corruption during the conversion. The integrity of health data cannot be guaranteed each time it is ported into a new system because current methods always require some kind of intervention - mapping, transformations, tweaking, 'cleaning', etc. Small errors can creep in with each data manipulation and which over time, can compromise the safety and value of our health data. On principle we know that the data should not be manipulated, but being limited by our traditional approach to siloed EHR applications, we have previously had little choice.
 We need to change our approach and preserve the integrity of our health data at all costs. After all it is the only reason why we record any facts or activity in an electronic health record - so we can use the data for direct patient care; share & exchange the data; aggregate and analyse the data, and use the data as the basis for clinical decision support.
We need to change our approach and preserve the integrity of our health data at all costs. After all it is the only reason why we record any facts or activity in an electronic health record - so we can use the data for direct patient care; share & exchange the data; aggregate and analyse the data, and use the data as the basis for clinical decision support.
We should not be focused on the application alone.
Apps will come and go, but we want our health data to persist - accurate and safe for clinical use - beyond the life span of any one clinical software application.
I've said this before, but it's worth saying many times over:
It's. all. about. the. data.
One of the key benefits of the openEHR paradigm is that the data specifications (the archetypes) are defined independently of any specific clinical system or application; are based on an open EHR architecture specification; and are publicly available in repositories such as the Clinical Knowledge Manager. It means that any data that is captured according to the archetype specification is directly usable by any and all archetype-compliant systems. Plus the data is no longer hard-wired into a proprietary application so that it is orders of magnitude easier to accurately share or transfer health data than it has before.
Clinical system vendors that don't directly embrace the archetype-technology may still 'archetype-aware', and can choose to use the archetype specifications as a means to understand the meaning of existing archetyped data and integrate it appropriately into their systems. Similarly they can map from their non-openEHR systems to the archetype specifications as a standardised method for data export and exchange.
The openEHR paradigm enables potential for archetype-compliant systems to share the same archetyped data repository - along the lines of an Apple platform 'plug & play' approach, with applications being added, removed or updated to suit the needs of the end-users, while the data persists intact. No more data conversions needed.
Adapted from Martin van der Meer, 2009
Now that's good news for our health data.
Are we there yet?
No, but we are definitely moving in the right direction... Conversations are happening that were uncommon generally, and downright rare in the US only 18 months ago. I've been rabbiting on for some time about the need for a 'universal health record - an application-independent core of shared and standardised health information into which a variety of 'enlightened' applications can 'plug & play'; thus breaking down the hold of the proprietary and 'not invented here' approach of proprietary clinical applications with which we battle most everywhere today.
So it was pleasing to see Margalit Gur-Arie's recent blog post on Arguments for a Universal Health Record. While I'm not convinced about the reality a single database (see my comments at the end of Margalit's post), I wholeheartedly endorse the principle of having a single approach to defining the data - this is a very powerful concept, and one that may well become a pivotal enabler to health IT innovation.
In addition, Kevin Coonan has started blogging in recent days - see his Summary of DCMs regarding principles of Detailed Clinical Models (aka DCMs). Now I know that Kevin's vision for an implementable HL7 DCM is totally different to the openEHR DCMs (=archetypes) that I work with. But we do agree on the basic principles about the basic attributes of these models that he has outlined in his blog post - it is quite a good summary, please read it.
Now these two bloggers are US-based - and this is significant because in the US there has been a huge emphasis on connecting between systems and exchange of document-based health information up until recent times. I view their postings as indicative of a growing trend toward the realisation that standardisation of clinical content is a necessary component for a successful health IT ecosystem in the (medium-longterm, sooner the better) future.
Note that "Detailed Clinical Models", is the current buzz phrase for any kind of model that might be standardised and shared but is also used very specifically for the HL7 DCMs currently in the midst of an interminable ballot process and the Australian national program's DCMs, which are actually openEHR archetypes being used as part of their initial specification process. "Detailed Clinical Models" is being used in many conversations rather blithely and with many not fully understanding the issues. On one hand it is positively raising awareness of our need to standardise content and on the other hand, it is confusing the issue as there are so many approaches. See my previous post about DCMs - clarifying the confusion.
It is worth flagging that there has been considerable (and I would also venture to say, rather premature) effort put in by a few to formalise principles for DCMs in the draft ISO13972 standard (Quality Requirements and Methodology for Detailed Clinical Models), currently out for ballot. My problem with this ISO work is that the DCM environment is relatively immature - there are many possible candidates with as many different approaches. It is also important to make clear that having multiple DCMs compliant with generic principles outlined in an ISO standard may mean that the quality of our published silos of "DCM made by formalism X" and "DCM made by formalism Y" models might be of higher quality, but it definitely will not solve our interoperability issues. For that you need a common reference model underpinning the models or, alternatively, a primary reference model with known and validated transformations between clinical model formalisms.
The more recent evolution of the CIMI group is really important in this current environment. It largely shares the principles that Kevin, openEHR and ISO13972 espouse - creation of standardised and shareable clinical content models, bound sensibly to terminology, as the basis for interoperability. These CIMI models will be computable and human readable; they will be based on a single Reference Model (yet to be finalised) and common data types (also yet to be finalised), and utilising the openEHR Archetype Definition Language (ADL) 1.5 as its initial formalism. Transformations of the resulting clinical models to other formalisms will be a priority to make sure that all systems can consume these models in the future. All will be managed in a governed repository and likely under the auspice of some kind of an executive group with expert teams providing practical oversight and management of models and model content.
Watch for news of the CIMI group. It has a influential initial core membership that embraces multiple national eHealth programs and standards bodies, plus all the key players with clinical modelling expertise - bringing all the heavy lifters in the clinical modelling environment into the same room and thrashing out a common approach to semantic interoperability. They met for 3 days recently prior to the HL7 meeting in San Antonio. The intent (and challenge) is to get all of this diverse group singing from the same hymn book! I believe they are about to launch a public website to allow for transparency which has not been easy in these earliest days. I will post it here as soon as it is available.
Maybe the planets are finally aligning...!
I have observed a significant change in the mind sets, conversations and expectations in this clinical modelling environment, over the past 5 years, and especially in the past 18 months. I am encouraged.
And my final 2c worth: in my view, the CIMI experience should inform the ISO DCM draft standard, rather than progressing the draft document based on largely academic assumptions about clinician engagement, repository requirements and model governance - there is so much we still need to learn before we lock it into a standard. I fear that we have put the cart before the horse.
openEHR: interoperability or systems?
Thomas Beale (CTO of Ocean Informatics and chair of the Architecture Review Board of the openEHR Foundation) posted these two paragraphs as part of the background for his recent Woland's Cat post - The Null Flavour debate - part I. It is an important statement that I don't want to get lost amongst other discussion, so I've reposted it here:
An initial comment I will make is that there is a notion that openEHR is ‘about defining systems’ whereas HL7 ‘is about interoperability’. This is incorrect. openEHR is primarily about solving the information interoperability problem in health, and it addresses all information, regardless of whether it is inside a system or in a message. (It does define some reference model semantics specific to the notion of ‘storing information’, mainly around versioning and auditing, but this has nothing to do with the main interoperability emphasis.)
To see that openEHR is about generalised interoperability, all that is needed is to consider a lab archetype such as Lipid studies in the Clinical Knowledge Manager. This archetype defines a possible structure of a Lipids lab test result, in terms of basic information model primitives (Entry, History, Cluster, Element etc). In the openEHR approach, we use this same model as the formal definition of this kind of information is in a message travelling ‘between systems’, in a database or on the screen within a ‘system’. This is one of the great benefits of openEHR: messages are not done differently from everything else. Neither is interoperability of data in messages between systems different from that of data between applications or other parts of a ‘system’.
Anatomy of an archetype
 With this blog I want to establish a simple baseline statement or overview about openEHR archetypes, aggregated from other posts and publications – a reference point if you like – from which we can journey further and in more detail into the issues around clinical modelling using specific archetypes.
Archetypes are a strong basis for data liquidity. Create the archetype once, agree through peer-review, re-use when and where required – the foundation for a 'universal health record'.
With this blog I want to establish a simple baseline statement or overview about openEHR archetypes, aggregated from other posts and publications – a reference point if you like – from which we can journey further and in more detail into the issues around clinical modelling using specific archetypes.
Archetypes are a strong basis for data liquidity. Create the archetype once, agree through peer-review, re-use when and where required – the foundation for a 'universal health record'.
Definition:
Formal Definition
An archetype is a computable expression of a domain content model in the form of structured constraint statements, based on a reference (information) model. openEHR archetypes are based on the openEHR reference model. Archetypes are all expressed in the same formalism. In general, they are defined for wide re-use, however, they can be specialized to include local particularities. They can accommodate any number of natural languages and terminologies. [Archetype Definitions and Principles]
Clinician's 'lay' definition:
An archetype is a structured, computable specification for one single, discrete clinical concept. - much simpler!
Each archetype is a rich health data specification, defined and agreed by the clinicians themselves to ensure that each model is 'fit for clinical purpose'. Collectively these archetypes together can create an electronic health record 'lingua franca', or an example of PCAST report's 'Universal Exchange Language (although archetypes are not limited only to exchange of health information).
Archetypes can be used to model a range of clinical concepts:
- Simple and straightforward - such as 'Blood Pressure'; 'Body Weight'; or 'Symptom'; or
- Complex, compound concepts - such as 'Medication order', or recording how a 'Procedure' has been carried out.
Purpose
Archetypes are:
- Potential 'agents for change' – allowing knowledge-driven EHRs as an orthogonal approach to EHR development.
-
The basis for a coherent and consistent approach for the whole continuum of clinical care and related activities:
- Recording, storing and querying health information in electronic health information;
- Exchanging health information - the broader the agreement & governance, the greater the potential for interoperability;
- Data aggregation;
- Knowledge-based activities; and
- Comparative data analysis.
Using archetypes requires:
- A change of mindset – no longer silos of data; no longer message and document driven.
- Upfront planning & coordination in development of archetypes, but not necessarily more time
-
CLINICIAN LEADERSHIP & ENGAGEMENT
- To ensure the quality of clinical data in our EHRs
- To warrant the clinical data is safe & 'fit for purpose'
Design
Each archetype is designed:
- As a maximal data set effectively everything one can think of about a clinical concept in all situations by the consumer and any and all providers. This design intent is pragmatically constrained to be sensible in some situations, however good archetype design will always ensure that the full model can be achieved through nesting of additional archetypes to create the maximal dataset/universal use-case ideal.
- For the 'universal use-case' – for re-use in any and all scenarios, from use at home, to primary care, community care, hospital care, secondary use and for research.
Each archetype needs to represent the variety of data that is required to represent the information required for all aspects of clinical care and related activities, including:
- The ability to capture both free-text vs structured data;
- Normal statements such as "Nil significant" or "NAD";
- Graphable data;
- Images – eg a homunculus, or a surgical diagram;
- Multimedia including photos, video and audio or an ECG wave format;
- Questionnaires, checklists etc
Think about how the clinician may record the data: different clinicians will need differing approaches to recording their health information and the models need to allow for this clinical diversity. Some will prefer to use more free text and others more structure; some need more detail than others; we need to capture current requirements for recording and exchange while keeping in mind what might be best practice in the near and distant future.
The best designers, without a doubt, have a clinical background. I have seen clinicians and non-clinicians given identical clinical content, but the archetypes produces are surprisingly different. Clinician modellers definitely model their domain better than those who are not familiar with clinical practice. They are perhaps better able to envision the fractal nature of medicine and take that in to account in creating archetypes, especially the complex areas which require nesting of archetypes within each other to capture the depth and richness required in the models.
Integration with terminologies such as SNOMED, ICD or LOINC are strategic and important – the structure of an archetype is not enough to represent clinical information adequately, and similarly terminology alone is inadequate. However the combination of terminology within an archetype structure, by naming elements or provision of value sets, is semantically very powerful. There is still considerable work to be done to determine how to represent the information overlap – that which can be expressed either in the archetype structure itself or via terminology. This remains a 'grey zone' and is a pressing area for collaboration or research.
Archetype classes – support clinician's processes
There are four main categories of archetypes that are useful to understand – each corresponding to classes in the openEHR Reference Model.
1. Compositions – which correspond to commonly used clinical documents, such as 'antenatal visit' or 'care plan'. 2. Sections – these are effectively used to assist with human navigation within EHRs and correspond to document headings, for example 'antenatal examination' or 'summary'.
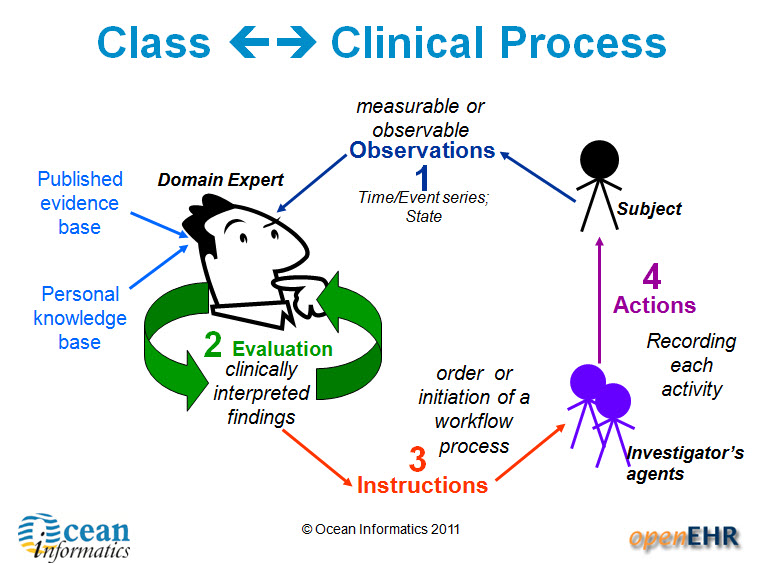
3. Entries – these are the most common and are fundamental building blocks of EHRs. There are four main types of Entry archetypes:
- Observations – recording measurable or observable data e.g. blood pressure, symptoms or weight;
- Evaluations – recording clinically interpreted findings e.g. adverse event, diagnosis or assessment of risk;
- Instructions – recording the initiation of a workflow process, such as a medication order or referral;
- Actions – recording clinical activities e.g. procedure or medication administration. Actions complement the instruction and can record the ensuing state of the instruction, such as 'completed' or 'cancelled'.
4. Clusters
Lifecycle & Governance
The Clinical Knowledge Manager (CKM) – www.openEHR.org/knowledge - is an international, online clinical knowledge repository under the auspice of the openEHR Foundation that provides:
- Access to a library of cohesive archetypes and related knowledge artefacts;
- Peer review, life cycle management & publication process via a Web 2.0 approach for clinical content, terminology binding, terminology reference set binding and translations;
-
Clinical Knowledge governance underpinned by a digital asset management system and providing:
- Provenance, audit trails, validation checking
- Release sets for implementation
Archetypes are freely available under a Creative Commons license.
Use
The need for EHRs to be able to represent clinical diversity is absolutely underrated. Existing EHR systems corral clinicians into certain types of recording etc, but for safety we need more diverse ways for clinicians to record what they need for patient care, and also to ensure that what is exchanged is appropriate for the patient and their individual circumstances. A 'one size fits all' document approach to emergency summaries or messages can be potentially unsafe. Elderly, pregnancy, chronic disease, paediatrics are extremely common examples where there are specific additional requirements needed to summary data sets
Templates are computable specifications for a specific clinical scenario or use-case. The openEHR paradigm takes the governed and agreed archetypes from CKM and creates clinically useful specifications that can be implemented in systems by:
- Aggregating the necessary archetypes together; and
- Constraining the maximal dataset archetypes so that only relevant data elements are active.
Examples include all clinical content required for a specific Message, Document, Clinical consultation or Report e.g. a histopathology report, a referral, an antenatal visit or a discharge summary.
In this way we can, in colloquial terms, 'have our cake and eat it too'. At the same time, CKM ensures tight clinical knowledge governance, yet templates allow for expression of clinician diversity.
Benefits
Archetypes provide a 'glide path' to knowledge-level interoperability of health information. They can:
- Bootstrap new application development – a source of agreed clinical content, avoid re-inventing the wheel for each vendor, organisation or project;
- Provide a forward 'roadmap' for existing applications – supporting gradual transition from proprietary silos to common data representations, perhaps via mapping to common messages as an interim step;
- Support data integration – the means to migrate valuable silos of legacy data to a common, re-usable representation
- Enable data aggregation & comparative analysis – the basis for ensuring 'apples' are 'apples', not 'oranges';
- Simplify messaging – keep message content consistent with EHR content;
- Enable knowledge-based activities eg Clinical Decision Support Systems – consistent & coherent; create once, re-use in all systems.
Don’t re-invent the (clinical content) wheel...
 It was with great interest that I read about the the recommendation for a universal exchange language in the recent release of the US report to the President: REALIZING THE FULL POTENTIAL OF HEALTH INFORMATION TECHNOLOGY TO IMPROVE HEALTHCARE FOR AMERICANS: THE PATH FORWARD.
I had asked the Direct project about the existence of a national plan for standardising clinical content only recently... It appeared that here was a plan after all.
It was with great interest that I read about the the recommendation for a universal exchange language in the recent release of the US report to the President: REALIZING THE FULL POTENTIAL OF HEALTH INFORMATION TECHNOLOGY TO IMPROVE HEALTHCARE FOR AMERICANS: THE PATH FORWARD.
I had asked the Direct project about the existence of a national plan for standardising clinical content only recently... It appeared that here was a plan after all.
So, to the report. The approach and benefits proposed started well...
The best way to achieve a national health IT ecosystem is to ensure that all electronic health systems can exchange data in a universal exchange language. The systems themselves could be designed in any manner desired — they could accommodate legacy systems that prevail or new recordkeeping systems and formats. The only requirement would be that the systems be able to send and receive data in the universal exchange language. (p41)
I have previously blogged about a universal health record underpinned by an application independent library of clinical content definitions, so the intent and benefits are well aligned with my preferred approach.
But then alarm bells started to ring....
Because of its multiple advantages, we advocate a universal exchange mechanism for health IT that is based on tagged data elements in an extensible markup language. If there were another equally good solution, it should also be considered; we have collectively been unable to think of one. (p43)
Issue #1: Isn't it more appropriate for step one to identify the need for standardised clinical content as a policy, rather than specify the format up front? Isn't that really the domain of health informatics experts as part of a subsequent work plan? I feel like we've skipped a couple of steps in the decision-making process. And are they really advocating the creation of this metadata-tagged XML from a zero starting point?
Issue #2: The last 9 words of that paragraph, "...we have collectively been unable to think of one." I'm glad that they are still open to equally good solutions being considered as indeed there are many ways that individuals, groups and organisations are exploring how to standardise clinical content definitions as the basis for a universal exchange mechanism.
In ISO TC 215, the International Standards Organisations Technical Committee for Health Informatics, there is a new work item which has been evolving for at least 2 years, although yet to attain committee draft status, known as ISO 13972 - Quality criteria for detailed clinical models. This work item is targeting a new international standard for determining quality criteria about the development of detailed clinical models - all clinical models, pick your flavour! In the world of international standards it has been recognised for years that with the plethora of different approaches to developing clinical models for EHRs, there is a need for some criteria to support quality aspect in their development. This work is being led by modellers from the Netherlands, with experts participating from the Australian, Danish, German, Swedish, US and Canadian standards organisations. Creating clinical content is definitely not a new field of endeavour by the time it enters the international standards arena.
So, I am extremely surprised that this expert PCAST group have not been able to 'collectively think' of an existing alternative.
In my last blog - Clinical Knowledge Governance in a Web 2.0 world – I pointed to a number of approaches to standardised clinical content to support health information exchange.
1. In the US – including, but by no means limited to:
- the HL7 standards organisation - where my UK colleague, Charlie McKay, informs me that there are more than 20 different approaches to clinical content development. Keith Boone (@motorcycle_guy) has posted his response to the PCAST report from a HL7 point of view - The Language of HealthIT;
- Stan Huff's group at Intermountain Health in Utah have had extensive experience in defining standardised clinical content across all of Intermountain's systems – they are leading experts in this domain; and
- I understand Don Mon and his team from AHIMA have also been working in this area.
2. In Europe, and Australia:
- The Australian National eHealth Transition Authority (NEHTA) has just launched a national clinician-led CKM approach to grassroots clinical content standardisation creating NEHTA's Detailed Clinical Models (DCMs).
- The Swedish national eHealth program has been actively working to integrate openEHR archetypes with its' VTIM architecture.
In addition, a few more points...
Firstly, the focus of the PCAST report is still only on data exchange, not on ensuring a sound foundation of a person-centric electronic health record. I'll say it again... get the data right and then the data will be able to be re-used, to multitask, be liquid, flowing to where it needs to be. It will become the solid foundation on which to build lifelong health records, simpler health information exchange, data integration & aggregation, research, reporting and knowledge-based activities. By focusing on exchange alone, then... you'll hopefully be able to exchange well and the rest will be considerably more uncertain.
Secondly, the proposed variant of XML is described as a 'straightforward' and 'superior' solution (p44), and the assumption that it will be scalable, protected by encryption, and that data element access services will be enough to support the health information exchange required. By contrast HL7, ISO/CEN 13606 and openEHR have taken decades to develop and refine underlying reference models to ensure that they have an unambiguous, consistent, secure way to represent personal health information – so you know who created the data, who is the subject of care, what the data means, what are the access rules applicable etc. In the openEHR environment, the specification authors developed Archetype Definition Language (ADL) for the purpose - and now part of the ISO 13606 standard - because the alternatives such as standard XML were not robust enough to represent health information. A 'straightforward' XML approach has a strong possibility of failure without a RM underpinning it.
And finally, there is the area of clinical knowledge governance itself. Health is dynamic, complex and diverse. The work required to represent healthcare as computable clinical content definitions or specifications is huge – don't underestimate the sheer volume of work that will be required. It is not realistic to expect a 'rapid mapping' of existing proprietary data structures into tagged data elements. Who will decide the clinical content in the models? If there are over 7000 clinical vendors in the US, which will be 'the source' or sources? Which are 'correct' or 'authoritative'? What methodology will be used to create the models? What level of granularity for each clinical element? How will they be aggregated together to represent clinical documents or events, and constrained to be useful for the clinical purpose? I have a million more questions...
Once the information models are defined, there will be a need for them to be validated before they can become the basis for a standardised or national clinical content library – suitable for consumers, clinicians, organisations, vendors, researchers and jurisdictions. A requirement will be recognised for life-cycle management and publication of these models, roadmaps for legacy data to migrate towards, and harmonise with, the new national health information 'source of truth', plus ongoing maintenance and governance.
Eric Browne stated in his recent blog, Recasting e-Health in the USA:
The work in Sweden, the UK, Singapore and even Australia, based on openEHR or ISO 13606 archetypes (i.e. implementable renditions of Detailed Clinical Models) is far more advanced and promising than that offered by the PCAST approach.
openEHR, which is my interest, has an approach to defining, agreeing and governing clinical content models for electronic health records, known as archetypes. It has taken more than 18 years to develop the openEHR technical specifications, and the last 10 years to achieve its' current approach and position in terms of clinical modelling. It is gaining traction, albeit with a modest volunteer community, especially now that it has a collaborative portal, known as the Clinical Knowledge Manager, to support sharing or models, reviews of clinical content, translation and terminology binding, and model governance.
Standardising health information definitions for health records or exchange is not a trivial task. Learn from what has already been achieved – all shapes, flavours and doctrines. Whatever you do, don't reinvent the wheel and create yet another universal language!
Clinician-led eHealth records - a knowledge-enabled approach
My presentation given to W.H.O. in Geneva last week... [slideshare id=5492832&doc=20101008whoclinician-ledehealthrecords-101019133425-phpapp02]
What is the vision for an Open Health platform?
 Came across a tweet this morning from Tim O'Reilly (@timoreilly) linking to an article by Mark Drapeau (@cheekygeeky) entitled 'What is the Vision for Open Government Entrepreneurship'.
The first paragraph in particular caught my eye:
Came across a tweet this morning from Tim O'Reilly (@timoreilly) linking to an article by Mark Drapeau (@cheekygeeky) entitled 'What is the Vision for Open Government Entrepreneurship'.
The first paragraph in particular caught my eye:
Tim O’Reilly often explains Open Government, or Government 2.0, as “Government as a Platform” on which citizens build things for each other and participate in their government (rather than treating it like a vending machine). The co-founder of Personal Democracy Forum and techPresident Andrew Rasiej has a similar notion that he terms WeGovernment.
And then following the link to Tim O'Reilly's 2009 article, “Government as a Platform”:
...But as with Web 2.0, the real secret of success in Government 2.0 is thinking about government as a platform
. If there’s one thing we learn from the technology industry, it’s that every big winner has been a platform company: someone whose success has enabled others, who’ve built on their work and multiplied its impact. Microsoft put “a PC on every desk and in every home,” the internet connected those PCs, Google enabled a generation of ad-supported startups, Apple turned the phone market upside down by letting developers loose to invent applications no phone company would ever have thought of. In each case, the platform provider raised the bar, and created opportunities for others to exploit.
There are signs that government is starting to adopt this kind of platform thinking.
It got me wondering... When will the eHealth community start thinking in these terms?
- Open Government; Open Health.
- Government as a platform; Health as a platform.
- A cohesive platform approach rather than fragmented proprietary silos.
- Opportunities and innovations as spin-offs.
I wrote in a previous post about the concept of a universal health record based on an open, standardised architecture as the basis for a cohesive and sustainable approach to recording and exchanging health information, health data aggregation, support for knowledge-based activities such as clinical decision support and comparative data analysis. I also posted, rather naively perhaps, about the openEHR platform in which I work, as an open source health equivalent of the iPod/IPhone platform.
I’m sure most won’t accept a Microsoft-, Google- or Apple-equivalent as the platform and will push for an open option. And whatever you do, don't confuse open source software applications with the concept of an open source platform. An open platform can enable all software developers to play, no matter what their philosophy - 'what is under the hood' is open source and standardised - and that is a key differentiator.
But you get the idea, I think - the underlying notion of an Open Health platform is sound. Other knowledge domains are embracing the concept and way ahead in terms of innovation in this space.
When will the health domain start to engage in the same open-minded and innovative manner? Only then can we start to think in terms of Open Health Entrepreneurship as mentioned in @cheekygeeky's article.
Time for a revolution, me thinks!
Archetypes: the ‘glide path’ to knowledge-enabled interoperability
 In a world where connectivity is the universal aspiration, our health information is largely still caught up in silos and, in the main, is not accessible to those who need it – patients, clinicians, researchers, epidemiologists and planners. Shared electronic health records (EHRs) are increasingly needed to support the improvement of health outcomes by providing a timely, comprehensive and coordinated foundation for provision of healthcare. For decades people have been attempting to share health information, but the incremental approach has not been wholly successful – progress has been made, but despite enormous investment and resources, the solution has been found to be more difficult than most anticipated; many well-funded attempts have been stunningly unsuccessful. Healthcare provision appears not to fit the model that has been so successful in other domains such as banking or financial services. Why has sharing health information been so difficult? After all, on the surface, data are, simply, data.
In a world where connectivity is the universal aspiration, our health information is largely still caught up in silos and, in the main, is not accessible to those who need it – patients, clinicians, researchers, epidemiologists and planners. Shared electronic health records (EHRs) are increasingly needed to support the improvement of health outcomes by providing a timely, comprehensive and coordinated foundation for provision of healthcare. For decades people have been attempting to share health information, but the incremental approach has not been wholly successful – progress has been made, but despite enormous investment and resources, the solution has been found to be more difficult than most anticipated; many well-funded attempts have been stunningly unsuccessful. Healthcare provision appears not to fit the model that has been so successful in other domains such as banking or financial services. Why has sharing health information been so difficult? After all, on the surface, data are, simply, data.
Why is the health information domain different?
Health information is the most multifaceted and largest knowledge domains to try to represent in a computer. The SNOMED CT terminology alone has over 450,000 terms expressing health-related concepts, and our collective knowledge about health is far broader, deeper and richer than that required to represent financial systems. The added bonus in health is that our information domain is dynamic - growing and changing as our understanding increases.
Recording, communicating and making sense of health information is something that clinicians do remarkably well in a localised, non-digital world. However the human cognitive processes and assumptions that underpin the traditional health records do not easily translate into the computerised environment. Consider the need for narrative versus structured data; the complexity of clinical statements; use of the same data in a variety of clinical contexts; the need for clinicians to make ‘normal’ or ‘nil significant’ statements, and also the oft underrated positive statements of absence; and the need for graphs, images or multimedia in a good health record. Grassroots clinicians have different personal preferences for creating their clinical records and to support their requirements for direct provision of clinical care and communication to colleagues. In parallel, jurisdictions have different expectations of the grassroots clinical data collection that will support reporting, data aggregation and secondary use of data.
Throw into this mix the complex and convoluted processes required to support healthcare provision; mobile patient populations; and the need for lifelong health records, and it starts to become easier to understand why eHealth has been more of a challenge that many first thought.
Information-driven EHRs
Traditional approaches to the development of EHRs have been software application-driven, hard-coding clinical knowledge into the proprietary data model for each software system and resulting in silos of health information locked away in proprietary databases. This is valuable data, and even more valuable if we can get access to it, exchange it and utilise it. Key stakeholders – patients, clinicians, researchers, planners and jurisdictions – are currently disempowered and are not easily able influence or express their data requirements. We have mistaken the software application for the electronic health record - a classic example of the ‘tail wagging the dog’.
If we focus on the electronic health record being the data, we turn the traditional paradigm upside down. Our EHRs become information-driven by putting the stakeholders at the centre to direct the information content and quality aspects of our EHR systems. It is only then that our systems will be able to reflect the real requirements of stakeholders, ensuring that health information collected data is ‘fit for use’ and will support personal health records, clinician health records and, with appropriate authorisation and permissions, the broadest range of secondary use.
Sharing health information requires common and coherent health information definitions or models – ensuring that health information can be expressed in a way that is meaningful to stakeholders AND that computers can process it. According to Walker et al , Level 4 interoperability, or ‘machine interpretable data’, comprises both structured messages and standardised content/coded data. In practice, it means that data can be transmitted and viewed by clinical systems without need for further interpretation or translation. This semantic, or knowledge-level, interoperability is absolutely required for truly shareable health records, data aggregation, knowledge-based activities such as clinical decision support, and to support comparative analysis of health data. Further, it is only when this health information model is agreed at a local, regional, national or international level, that true semantic interoperability can occur at each of these levels. The broader the level of clinical content model agreement, the broader the potential for semantic health information exchange.
The openEHR paradigm
openEHR is a purpose-built, open source, information-driven electronic health record architecture focused on ensuring that the grassroots health information is recorded clearly, coherently and unambiguously in EHRs, and supporting re-use in other contexts where appropriate. It adopts an orthogonal approach to EHRs - a dual-level modelling methodology with clear separation of the technical from the clinical domains, where software engineers focus on their application development and the clinical domain experts focus on the health information definitions. openEHR focuses on the data - using computable knowledge artefacts known as archetypes and templates to formally express health information.
openEHR archetypes are computable definitions created by the clinical domain experts for each single discrete clinical concept – a maximal (rather than minimum) data-set designed for all use-cases and all stakeholders. For example, one archetype can describe all data, methods and situations required to capture a blood sugar measurement from a glucometer at home, during a clinical consultation, or when having a glucose tolerance test or challenge at the laboratory. Other archetypes enable us to record the details about a diagnosis or to order a medication. Each archetype is built to a ‘design once, re-use over and over again’ principle and, most important, the archetype outputs are structured and fully computable representations of the health information. They can be linked to clinical terminologies such as SNOMED-CT, allowing clinicians to document the health information unambiguously to support direct patient care. The maximal data-set notion underpinning archetypes ensures that data conforming to an archetype can be re-used in all related use-cases – from direct provision of clinical care through to a range of secondary uses.
Templates are used in openEHR to aggregate all the archetypes that are required for a particular clinical scenario – for example a consultation or a report. These can also be shared, preventing more ‘wheel re-invention’. Individual content elements of each maximal archetype can be ‘disabled’ in the template so that the only data elements presented to the clinician are those that conform to national or local requirements and are relevant and appropriate for that use-case scenario. For example, a typical Discharge Summary may commonly comprise 10 common archetypes; templates allow the orthopaedic surgeon to express a slightly different ‘flavour’ of the Discharge Summary based on which elements of each archetype being either active or disabled, compared to that required by a Obstetrician who needs to share information about both mother and newborn. One ‘size’, or document, does not fit all. The archetypes, as building blocks, are the key to semantic interoperability; while templates allow flexible expression of the archetypes to fulfill use-case requirements.
How achievable is this? Only ten archetypes are needed to share core clinical information that could save a life in an emergency or provide the majority of content for a discharge summary or a referral. If each archetype takes an average of six review rounds to reach clinician consensus and each review round is open for 2 weeks, it is possible to obtain consensus within an average of three months per archetype – some complex or abstract ones may be longer; other simpler, more concrete archetypes will be shorter. Many archetypes are already well developed in the international arena and within national programs. As archetype reviews can be run in parallel, a willing community of clinicians could achieve consensus for core clinical EHR content within three to six months.
It is estimated that as few as fifty archetypes will comprise the core clinical content for a primary care EHR, and maybe only up to two thousand archetypes for a hospital EHR system including many clinical specialties. The initial core clinical content will be common to all clinical disciplines and can be re-used by other specialist colleges and interested groups. More specialised archetypes will gradually and progressively be added to enhance the core archetype pool over time.
The openEHR Clinical Knowledge Manager (CKM) is an online clinical knowledge management tool – www.openEHR.org/knowledge - which provides a repository for archetypes and other clinical knowledge artefacts, such as terminology subsets and document templates. Based on a data asset management platform it provides a clinical knowledge ecosystem supporting the publication lifecycle and governance of the archetypes. Within CKM, a community of grassroots clinicians and health informaticians collaborate in online reviews of each archetype until consensus is reached and the agreed archetype content is published. Clinicians and other domain experts need no technical knowledge to engage with archetypes - the technical aspects of archetypes are kept hidden ‘under the bonnet’ – but they use their expertise to ensure that the content definitions within each archetype is correct and appropriate. Each content review is conducted online at a time of convenience to the clinician and usually only takes five to ten minutes for each participant. Thus the clinical domain experts themselves drive the archetype content definitions, and CKM has become a peer-reviewed knowledge resource for all parties seeking shared, standardised and computable health information models.
At the time of writing CKM has acquired, largely by word of mouth, 565 registered users from 62 countries, including 181 people who have volunteered to review archetypes, and 73 who have volunteered to translate archetypes. The repository contains 273 archetypes, of which 15 have content that are in team review and 9 published. Two example templates have been uploaded, and we await final publication of the openEHR template specification before we expect to see template activity increase. Terminology subset functionality has been added only recently and our first terminology subsets uploaded. So, while CKM is still relatively new, its Web2.0 approach to artifact collaboration and publishing, combined with formal knowledge artifact governance positions CKM as a pioneering ‘one stop shop’ for clinical knowledge resources online.
Current CKM functionality includes:
- Display of artefacts including structured views, technical representations and mind maps to make it easy for clinicians and others to review;
- Uploading of new knowledge artefacts – archetypes, templates and terminology subsets – for review and publication;
- Archetype metadata supporting classification, ontological relationships and repository-wide searches;
- Digital asset management including provenance and artefact audit trail;
- Integration with openEHR tools supporting quality assessment & technical validation checks;
- Review and publication process for clinical content – draft, team review, published and reassess states
- Terminology binding and terminology subset reviews;
- Online archetype translation with review;
- Community engagement via threaded discussions, repository downloads, attached resources, watch lists, email notifications, user dashboards and release sets;,
- Editorial support via To Do lists, user and team administration, review management, artefact modification, classification management, broadcast emails etc;
- Subscriber auto-notification including Twitter and email
- Reports – Archetypes, Templates and Registered users
A governed repository of shared and agreed archetypes will provide a ‘glide path’ towards full semantic interoperability of health information; a clear forward path for standardisation of data definitions. These will bootstrap new application or program development, provide a ‘road map’ to support gradual transition of existing systems to common data representation and provide the means to integrate valuable silos of legacy data.
Benefits of a collaborative, data-driven approach
A collaborative and domain expert-led approach to our health information provides many benefits which include the following.
Benefits for stakeholders
- Active involvement of domain experts to ensure the safety and quality of health information.
- Development of a coherent set of health information definitions:
- Improved data quality – shared core clinical content plus specialised domain-specific content will be agreed and ratified by the domain expert community; health information created will need to conform to the agreed archetype specifications.
- Improved data ‘liquidity’ – specifications to support exchange, flow and re-use of health information - from direct patient care through to secondary use of data. Improved data longevity – shared non-proprietary health information definitions minimise need for data transformations or system migration and the inherent risk of data loss; will support the cumulative, lifelong health records and longitudinal data repositories;
- Improved data availability – easier integration of health information from disparate sources when based on common archetype definitions;
- Re-use, integrate and aggregate data for supporting quality processes such as clinical audit, reporting and research; and
- Break down the existing ‘silos’ of health information based on proprietary and varied definitions.
- Online collaboration maximises the potential for a breadth of grassroots stakeholder engagement in ensuring correctness of the health information definitions.
- Active participation by clinical domain experts to shape and influence their EHRs, ensuring that EHR content is ‘fit for clinical purpose’.
- Online participation in clinical content review will be of short duration and at times of convenience to the clinician, avoiding the significant time and opportunity cost of attendance at face-to-face meetings.
Benefits for patients
- Data created and stored in a shared, standardised and non-proprietary representation supports the potential for application-independent data records that can persist for the life of the patient.
- Improved data ‘liquidity’ – so that data can flow between healthcare providers and systems to where the patient needs it.
Benefits for national programs and other jurisdictions
- Development of a coherent national set of clinical content specifications to support the shared EHR programs, health information exchange and secondary use.
- Enables national governance of foundation clinical content while at the same time facilitates flexible expression of local domain requirements
- Efficient use of sparse clinical, informatics and stakeholder resources:
- Design & create an archetype once; re-use many times;
- Leveraging existing clinical specification work done internationally to improve local national pool of archetypes;
- Online collaboration maximise the potential for stakeholder engagement at the same time as minimising the requirement for expensive face-to-face meetings; and
- Review and publication of agreed clinical specification definitions within weeks to months;
- Review and standardisation of clinical documents containing agreed archetypes will be relatively short.
- • Clinical knowledge management ecosystem:
- Single national repository of clinical knowledge artefacts, including archetypes and terminology subsets.
- Focussed and coordinated knowledge management environment where all stakeholders can observe, participate and benefit; the opposite of the current fragmented, isolated and proprietary approach to defining health information content.
- Digital knowledge asset management:
- Manages authoring, reviewing, publication and update lifecycle of all knowledge assets;
- Provenance and asset audit trails;
- Ensures asset compliance to quality criteria;
- Ensures technical validation of assets; and
- Development of coherent release sets for implementers;
- Governance of knowledge assets.
- Distribution of knowledge assets via coherent release sets.
- Removes the need for per message or per document negotiation between application developers, organisations and jurisdictions each time information needs to be integrated or exchanged by use of the standardised content within more generic message wrappers or document structures.
- Transparency of editorial and publishing processes; accountability to the domain expert community itself.
- Precludes the need for ratification of clinical or reporting documents through a traditional standards process when they consist of subsets of the nationally agreed archetypes.
Benefits for application developers
- Download coherent sets of clinical content definitions from a published and agreed national repository – not re-inventing the wheel by defining each piece of health information over and over.
- Software development remains focused within the expert technical domain – user interface; workflow processes; security, data capture, storage, retrieval and querying; etc.
- Removes the need for per message or per document negotiation between vendors, organisations and jurisdictions each time information needs to be integrated or exchanged by use of the standardised content within more generic message wrappers or document structures.
Benefits for secondary users of data
- Existing data can be mapped to archetypes once only, and transformed into a validated and consistent format; new data can be captured and aggregated according to the same national archetype definitions.
- Data stored in a common representation can be more easily aggregated and integrated.
- Access to valuable legacy data that would otherwise be unavailable.
Agreed and shared representations of the health information, embracing existing stakeholder requirements and developed rapidly by an active online community, will kick-start and accelerate many currently fragmented eHealth activities. Sharing archetypes as the definition of our health information will not only provide a common basis for recording and exchanging health information but also simplify data aggregation of data, support knowledge-based activities and comparative data analysis. Perhaps even more compelling, we are making certain that our domain experts warrant that the data within our EHRs, and flowing between stakeholders, is safe and 'fit for purpose'.
Gimme an… uhr?
 What do we have?
You can have all the messages, connect-a-thons and profiles that you like - certainly you will achieve short- to medium-term connectivity wins but it is not enough. It is not sustainable and clinicians want and need to share relevant data when it is needed. Predetermined messages and documents will be a great start, but it's not enough.
What do we have?
You can have all the messages, connect-a-thons and profiles that you like - certainly you will achieve short- to medium-term connectivity wins but it is not enough. It is not sustainable and clinicians want and need to share relevant data when it is needed. Predetermined messages and documents will be a great start, but it's not enough.
You can invest huge resources in terminology - we definitely need it - but terminology in isolation is also not enough.
You can demand all the 'damn data' that you want, but non-standardised data will still largely not be computable - largely limited to read-only access.
What do we want?
We hear talk about:
- shared electronic health records;
- semantic interoperability; and
- health data liquidity.
Essentially these phrases describe different aspects of the same 'elephant in the room'; just using different jargon...
We expect our healthcare professionals to be able to safely exchange important health information. To have the right health information available at the right place at the right time. To prevent us repeating our health histories ad nauseum. In fact a huge percentage of patients assume that we can already do this! But we can't... yet. Shared health records are not easy.
What do we need?
A single data-driven "universal health record", an uhr! And please note the deliberate lack of capitals.
What on earth is an uhr, I hear you quite reasonably ask...? A uhr is simply a non-proprietary pool of standardised health data which can be used for any purpose we want - PHR, EHR, EMR, SEHR, IEHR, research, epidemiology, reporting & stats, shared care, clinical decision support & more. There is no particular EHR application associated with it; in fact it works with any and every software program, vendor and healthcare provider.
The key (and I say it yet again)... it's. all. about. the. data.
In addition, and supported by:
- Collaboration at international, national, jurisdictional or organisation level to share and re-use the clinical data definitions, so that our health information can similarly be shared.
- Clinicians driving the clinical content definitions - reflecting the information they need for recording health care for themselves and their patients, ensuring that EHRs are 'fit for purpose'. Terminology integrated with these structured clinical content definitions to unambiguously define the building blocks of the health record.
As a result:
- Message structures required for data exchange become more about the wrapper and less about content - much easier to negotiate and maintain.
- Vendors no longer have clinicians locked-in to proprietary data silos
- Tools used to do 'clever stuff' with our data need only to be built once to conform to these agreed clinical data definitions - think decision support, reporting, data repositories, and integration. Now that takes away a lot of the barriers we strive to overcome!
- Data in a common format can be exchanged between systems - data liquidity becomes a reality, flowing to where it is needed and according to authorisation and access permissions.
Gimme a universal health record. Gimme my damn, um,... uhr, and let my data flow...
And so, I archetype;-)